The optimizeR package provides an object-oriented framework for optimizer functions in R and adds convenience features for minimization and maximization.
You probably do not need this package if you…
- already know which optimizer you want to use and are comfortable with its constraints, such as minimization only over the first function argument;
- want to compare optimizers that are already covered by
{optimx}, a framework for comparing about 30 optimizers; - are searching for new optimization algorithms, because this package does not implement optimizer functions itself.
You might find the package useful if you want to…
- compare optimizer functions not covered by optimx or other frameworks (see the CRAN Task View: Optimization and Mathematical Programming for an overview);
- have consistently named inputs and outputs across different optimizers;
- view optimizers as objects, which can help when implementing packages that depend on optimization;
- use optimizers for both minimization and maximization,
- optimize over more than one function argument,
- measure computation time or set a time limit for long optimization tasks.
How to use the package?
The following demo is a bit artificial, but it showcases the package purpose. Assume we want to
- maximize a function over two of its arguments,
- interrupt optimization if it exceeds 10 seconds,
- compare the performance of
stats::nlmandpracma::nelder_mead.
We can do this task with optimizeR. You can install the released package version from CRAN with:
install.packages("optimizeR")Then load the package via library("optimizeR").
1. Define the objective function
Let with
f <- function(a, b, x, y) {
a * exp(-0.2 * sqrt(0.5 * (x^2 + y^2))) + exp(0.5 * (cos(2 * pi * x) + cos(2 * pi * y))) - exp(1) - b
}For a = b = 20, this is the negative Ackley function with a global maximum in x = y = 0:
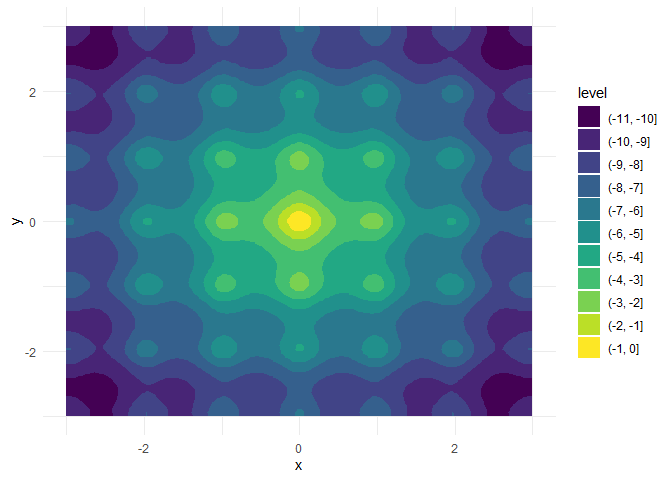
We want to keep a and b fixed here and optimize over x and y, which are both single numeric values.
Two problems would occur if we optimized f directly with, say, stats::nlm:
- There are two target arguments (
xandy). - The target arguments are not the first arguments of
f.
Both features are unsupported by stats::nlm and most other optimizers, but they are supported by optimizeR. We just have to define an objective object that we can later pass to the optimizers:
objective <- Objective$new(
f = f, # f is our objective function
target = c("x", "y"), # x and y are the target arguments
npar = c(1, 1), # the target arguments have both a length of 1
"a" = 20,
"b" = 20 # a and b have fixed values
)2. Create the optimizer objects
Now that we have defined the objective function, let’s define the optimizer objects. For stats::nlm, this is a one-liner:
nlm <- Optimizer$new(which = "stats::nlm")The optimizeR package provides a dictionary of optimizers that can be selected directly via the which argument. For an overview of available optimizers, see:
optimizer_dictionary
#> <Dictionary> optimization algorithms
#> Keys:
#> - lbfgsb3c::lbfgsb3c
#> - lbfgsb3c::lbfgsb3
#> - lbfgsb3c::lbfgsb3f
#> - lbfgsb3c::lbfgsb3x
#> - stats::nlm
#> - stats::nlminb
#> - stats::optim
#> - ucminf::ucminfAny optimizer that is not contained in the dictionary can still be put into the optimizeR framework by setting which = "custom" first:
nelder_mead <- Optimizer$new(which = "custom")
#> Use method `$definition()` next to define a custom optimizer.Then use the $definition() method:
nelder_mead$definition(
algorithm = pracma::nelder_mead, # the optimization function
arg_objective = "fn", # the objective function argument name
arg_initial = "x0", # the argument name for the initial values
out_value = "fmin", # the optimal function value element
out_parameter = "xmin", # the optimal parameter element
direction = "min" # the optimizer minimizes
)3. Set a time limit
Each optimizer object has a field called $seconds, which equals Inf by default. You can set a different single numeric value to define a time limit in seconds for the optimization:
nlm$seconds <- 10
nelder_mead$seconds <- 10Note that not everything, especially compiled C code, can technically be timed out. See the help page help("withTimeout", package = "R.utils") for more details.
4. Maximize the objective function
Each optimizer object has the methods $maximize() and $minimize() for function maximization and minimization, respectively. Both methods require values for two arguments:
-
objective(either an objective object as defined above or just a function) and -
initial(an initial parameter vector where the optimizer should start)
They optionally accept additional arguments for the objective function. Optimizer-specific arguments should be set when creating the optimizer object or via $set_arguments().
nlm$maximize(objective = objective, initial = c(3, 3))
#> $value
#> [1] -6.559645
#>
#> $parameter
#> [1] 1.974451 1.974451
#>
#> $seconds
#> [1] 0.04005599
#>
#> $initial
#> [1] 3 3
#>
#> $error
#> [1] FALSE
#>
#> $gradient
#> [1] 5.757896e-08 5.757896e-08
#>
#> $code
#> [1] 1
#>
#> $iterations
#> [1] 6
nelder_mead$maximize(objective = objective, initial = c(3, 3))
#> $value
#> [1] 0
#>
#> $parameter
#> [1] 0 0
#>
#> $seconds
#> [1] 0.02831006
#>
#> $initial
#> [1] 3 3
#>
#> $error
#> [1] FALSE
#>
#> $count
#> [1] 105
#>
#> $info
#> $info$solver
#> [1] "Nelder-Mead"
#>
#> $info$restarts
#> [1] 0Note that
the inputs for the objective function and initial parameter values are named consistently across optimizers,
the output values for the optimal parameter vector and maximum function value are also named consistently across optimizers,
the output contains the initial parameter values, the optimization time in seconds, and other optimizer-specific elements,
pracma::nelder_meadoutperformsstats::nlmhere both in optimization time and in convergence to the global maximum.
Getting in touch
If you have any questions, found a bug, or need a feature, please file an issue on GitHub.
